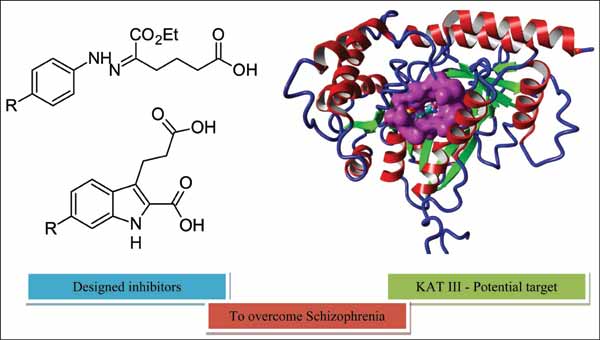
Abstract
Kynurenine aminotransferase (KAT) isozymes are responsible for catalyzing the conversion of kynurenine (KYN) to kynurenic acid (KYNA), which is considered to play a key role in central nervous system (CNS) disorders, including schizophrenia. The levels of KYNA in the postmortem prefrontal cortex and in the Cerebrospinal fluid (CSF) of schizophrenics are greater than normal brain. A basic strategy to decrease kynurenic acid levels is to promote the inhibition of the biosynthetic KAT isozymes. As there is no crystallographic model for human kynurenine aminotransferase III (KAT III), therefore, homology modeling has been performed based on the Mus musculus kynurenine aminotransferase III crystal structure (PDB ID: 3E2Y) as a template, and the model of the human KAT III was refined and optimized with molecular dynamics simulations. Further evaluation of the model quality was accomplished by investigating the interaction of KAT III inhibitors with the modeled enzyme. Such interactions were determined employing the AutoDock 4.2 program using the MGLTools 1.5.6 package. The most important interactions for the binding of the inhibitors, which are probably also central components of the active site of KAT III, were identified as Ala134, Tyr135, Lys 280, Lys 288, Thr285 and Arg429, which provide hydrogen bond interactions. Additionally, Tyr135 and Arg429 have good electrostatic interactions with inhibitors consistent with these residues also being essential for inhibition of the enzyme activity. We expect that this model and these docking data will be a useful resource for the rational design of novel drugs for treating neuropathologies.
Keywords: Docking, homology modeling, kynurenine aminotransferase, molecular dynamics simulation, schizophrenia.